
Main article: Protein Data Bank (file format)

Historically, the number of structures in the PDB has grown at an approximately exponential rate, with 100 registered structures in 1982, 1,000 structures in 1993, 10,000 in 1999, and 100,000 in 2014. The data of such structures is stored on the "electron density server". Clicking on the numbers in the linked external table displays examples of structures determined by that method.įor PDB structures determined by X-ray diffraction that have a structure factor file, their electron density map may be viewed. After 2013, a growing number of proteins are determined by cryo-electron microscopy. The final conformation of the protein is obtained from NMR by solving a distance geometry problem. When using X-ray diffraction, approximations of the coordinates of the atoms of the protein are obtained, whereas using NMR, the distance between pairs of atoms of the protein is estimated. Most structures are determined by X-ray diffraction, but about 10% of structures are determined by protein NMR. 4,718 structures in the PDB have a 3DEM map file deposited in EM Data Bank 4,814 structures in the PDB have a chemical shifts file. 10,289 structures have an NMR restraint file. As of 1 April 2020, the PDB comprised:ġ34,146 structures in the PDB have a structure factor file. The PDB database is updated weekly ( UTC+0 Wednesday), along with its holdings list. MX = macromolecular crystallography, 3DEM = 3D Electron Microscopy. Rate of Protein Structure Determination by Method and Year.
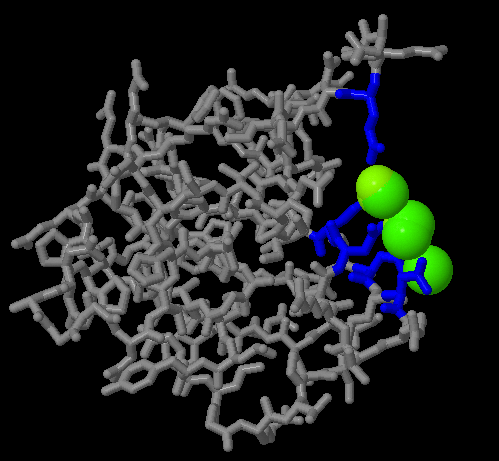
JMOL GFP PROTEIN DATABANK SOFTWARE
The data are then automatically checked for plausibility (the source code for this validation software has been made available to the public at no charge). The data processing refers to the fact that wwPDB staff review and annotate each submitted entry. Each of the four members of wwPDB can act as deposition, data processing and distribution centers for PDB data. The founding members are PDBe (Europe), RCSB (USA), and PDBj (Japan). In 2003, with the formation of the wwPDB, the PDB became an international organization. Berman of Rutgers University (one of the managing institutions of the RCSB, the other being the San Diego Supercomputer Center at UC San Diego). The PDB was transferred to the Research Collaboratory for Structural Bioinformatics (RCSB) the transfer was completed in June 1999. In January 1994, Joel Sussman of Israel's Weizmann Institute of Science was appointed head of the PDB. Upon Hamilton's death in 1973, Tom Koeztle took over direction of the PDB for the subsequent 20 years. The Protein Data Bank was announced in October 1971 in Nature New Biology as a joint venture between Cambridge Crystallographic Data Centre, UK and Brookhaven National Laboratory, US. SEARCH was instrumental in enabling networking, thus marking the functional beginning of the PDB. By 1971, one of Meyer's programs, SEARCH, enabled researchers to remotely access information from the database to study protein structures offline. In 1969, with the sponsorship of Walter Hamilton at the Brookhaven National Laboratory, Edgar Meyer ( Texas A&M University) began to write software to store atomic coordinate files in a common format to make them available for geometric and graphical evaluation. Two forces converged to initiate the PDB: a small but growing collection of sets of protein structure data determined by X-ray diffraction and the newly available (1968) molecular graphics display, the Brookhaven RAster Display (BRAD), to visualize these protein structures in 3-D. For example, SCOP and CATH classify protein structures, while PDBsum provides a graphic overview of PDB entries using information from other sources, such as Gene ontology. Many other databases use protein structures deposited in the PDB. Most major scientific journals and some funding agencies now require scientists to submit their structure data to the PDB. The PDB is a key in areas of structural biology, such as structural genomics. The PDB is overseen by an organization called the Worldwide Protein Data Bank, wwPDB. The data, typically obtained by X-ray crystallography, NMR spectroscopy, or, increasingly, cryo-electron microscopy, and submitted by biologists and biochemists from around the world, are freely accessible on the Internet via the websites of its member organisations (PDBe, PDBj, RCSB, and BMRB ). The Protein Data Bank ( PDB) is a database for the three-dimensional structural data of large biological molecules, such as proteins and nucleic acids.


 0 kommentar(er)
0 kommentar(er)
